aPROMISE
Introducing aPROMISE
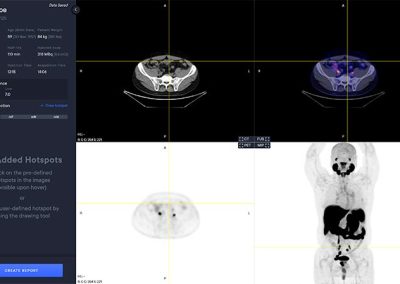
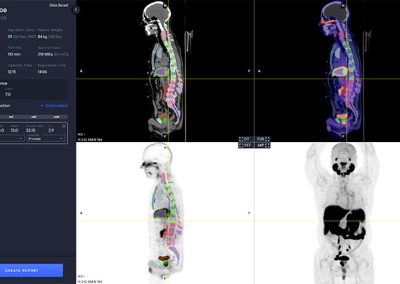
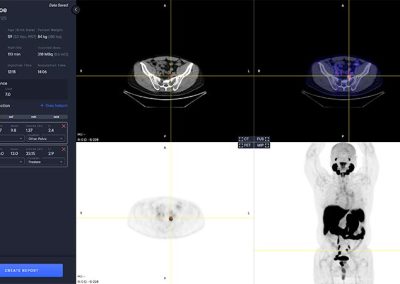
aPROMISE10 is a PACS platform that offers quantitative analysis and standardized reporting of PSMA PET/CT image assessments. Deep Learning technology on PSMA images to enhance:
- Efficiency: Reduce the laborious task of defining and locating the disease
- Consistency: Enhance the reproducibility and reliability among the readers
- Accuracy: Maintain the high diagnostic accuracy
- Standardization: Enable rapid detection and volumetric quantification of disease burden in PSMA images.
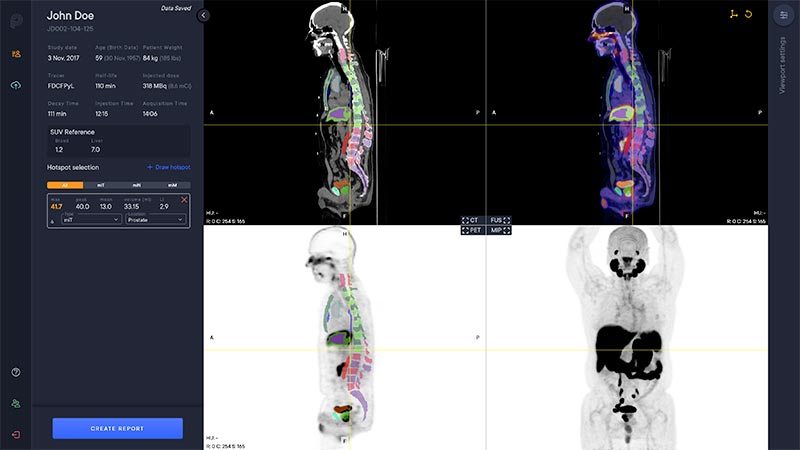
A deep learning-enabled application
- Automated segmentation and localization of PSMA PET lesion candidates
- Automated segmentation and quantification of PSMA uptake in reference organs
- Automated PSMA quantification at lesion level – tumor (miT), lymph node (miN, miMa), and visceral disease
Interested to learn more, book a demo!
Through rigorous performance studies, the aPROMISE has demonstrated:

Reproducible and Quantitative disease-burden Indices
- Improved inter-reader reproducibility in staging (κ >0.80) and quantification (ICC: 0.99) of prostate cancer patients2,3
- Quantitative PSMA scan index (PSI) was associated with PSA and Gleason Score2

Accurate Lesion Quantification
- High segmentation and detection accuracy (>90%) for PSMA lesions in regional and distant lymph Nodes.3

High Efficiency
- Significant efficiency in creating quantitative structure reporting. Reader spends an average ~3 min minutes for a comprehensive, quantitative report.2

Clinical Utility
- A quantitative PSMA score/index can help stratify patients for available treatment options2
- A quantitative PSMA score/index can accurately demonstrate disease progression and response2
Legal
Performance and Safety
isReferences
- Bubendorf, L., et al., Metastatic patterns of prostate cancer: an autopsy study of 1,589 patients. Human pathology, 2000. 31(5): p. 578-83.
- Pienta KJ, Gorin MA, Rowe SP, Carroll PR, Pouliot F, Probst S, et al. A Phase 2/3 Prospective Multicenter Study of the Diagnostic Accuracy of Prostate Specific Membrane Antigen PET/CT with (18)F-DCFPyL in Prostate Cancer Patients (OSPREY). The Journal of urology. 2021:101097JU0000000000001698.
- Morris MJ, Rowe SP, Gorin MA, Saperstein L, Pouliot F, Josephson D, et al. Diagnostic Performance of (18)F- DCFPyL-PET/CT in Men with Biochemically Recurrent Prostate Cancer: Results from the CONDOR Phase III, Multicenter Study. Clinical cancer research : an official journal of the American Association for Cancer Research. 2021.
- Eiber M, Herrmann K, Calais J, Hadaschik B, Giesel FL, Hartenbach M, et al. Prostate Cancer Molecular Imaging Standardized Evaluation (PROMISE): Proposed miTNM Classification for the Interpretation of PSMA-Ligand PET/CT. Journal of nuclear medicine : official publication, Society of Nuclear Medicine. 2018;59(3):469-78.
- Fanti S, Minozzi S, Morigi JJ, Giesel F, Ceci F, Uprimny C, et al. Development of standardized image interpretation for 68Ga-PSMA PET/CT to detect prostate cancer recurrent lesions. European journal of nuclear medicine and molecular imaging. 2017;44(10):1622-35.
- Rowe SP, Pienta KJ, Pomper MG, Gorin MA. PSMA-RADS Version 1.0: A Step Towards Standardizing the Interpretation and Reporting of PSMA-targeted PET Imaging Studies. European urology. 2018;73(4):485-7.
- Ceci F, Oprea-Lager DE, Emmett L, Adam JA, Bomanji J, Czernin J, et al. E-PSMA: the EANM standardized reporting guidelines v1.0 for PSMA-PET. European journal of nuclear medicine and molecular imaging. 2021;48(5):1626-38.
- Nickols N, Anand A, Johnsson K, et al. aPROMISE: A Novel Automated-PROMISE platform to Standardize Evaluation of Tumor Burden in (18)F-DCFPyL (PSMA) images of Veterans with Prostate Cancer. J Nucl Med. 2021.
- Johnsson K, Brynolfsson J, Sahlstedt H, et al. Analytical performance of aPROMISE: automated anatomic contextualization, detection, and quantification of [(18)F]DCFPyL (PSMA) imaging for standardized reporting. Eur J Nucl Med Mol Imaging. 2021.
- A unique configuration is also available in EU under the name PYLCLARI AI
GDPR
When you use aPROMISE to process patient information related to a patient who is a resident of the EU, you are responsible for ensuring that your organization complies with GDPR. In terms of GDPR you, as the user of aPROMISE, are the data controller and EXINI, as the service provider, is the data processor. In advance of processing data with aPROMISE, be sure that you have explicit consent from the patient whose data you are capturing. When data is sent to aPROMISE, it is stored in a secure manner, and is encrypted in transit and at rest.
Our Commitment
EXINI (‘we’ or ‘us’ or ‘our’) are committed and dedicated to ensuring the security and protection of the personal information that we process, and to provide a robust, continuous and consistent approach to data protection. Our objectives for GDPR and HIPAA compliance include the development and implementation of data protection roles, policies, procedures, controls and measures to ensure continuous safeguarding of the personal information under our remit.
How we are implementing GDPR and HIPAA
Policies & Procedures – Data protection policies and procedures to meet the requirements and standards of the GDPR and any relevant data protection laws, including HIPAA, are in place
Data Retention & Erasure – we have retention policies and are applying the privacy by design principle, meaning we store only data that is needed for the current task and only store it for as long as needed Data Breaches – as a medical device manufacturer we have breach procedures in place that ensure safeguards and measures to identify, assess, investigate and report any personal data breach at the earliest possible time
International Data Transfers & Third-Party Disclosures – when EXINI stores or transfers personal information outside the EU, we have robust procedures and safeguarding measures in place to secure,
encrypt and maintain the integrity of the data
Processor Agreements – when we use a third-party to process personal information on your behalf, we have data processor agreements and/or business associate agreements in place.