FDA cleared, software for quantitative analysis of PSMA PET/CT providing standardized, accurate, and rapid total-body quantitative output for PSMA PET/CT
PYLARIFY AI® provides accurate quantitative imaging biomarkers, increasing reproducibility in the assessment of disease2-6
Reach out and leave your information and one of our representatives will contact you!
Quantitative Imaging

HEALTHCARE PROFESSIONALS
Make more informed decisions on treatment selection, treatment eligibility, and monitoring of treatment response of prostate cancer patients

Supports longitudinal analysis
Supports longitudinal analysis of PSMA PET baseline and follow-up scans and allows clinicians to assess disease progression or monitor and follow response

Standardized report
- Generation of a standardized report harmonizes reads across multi disciplinary teams.
- In addition to quantitative biomarkers, the comprehensive report provides PSMA avidity at the lesion level that can inform eligibility of PSMA targeted therapies.
PYLARIFY AI® provides valuable imaging biomarkers that can be used to predict response to treatment and inform the optimal sequence of therapies in the metastatic setting, potentiating better patient outcomes1-10
How does PYLARIFY AI® work?
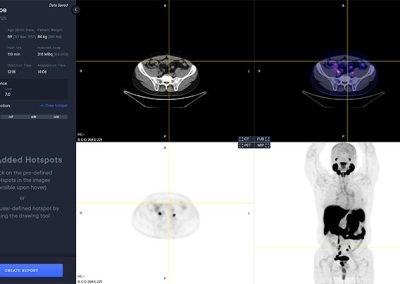
PYLARIFY AI® automatically analyzes the PET and CT image to detect target hotspots, regions of interest (ROIs) having locally elevated PSMA tracer intensities indicative of suspicious tumor tissue and metastasis. ROIs can be included in a report either by selecting them from the pre-segmented ones or by manually creating new ones. All pre-segmented ROIs are automatically quantified and labelled with volume, various SUV metrics as well as type, location and an index based on the promise criteria. Quantification of reference organs such as liver, parotid glands and blood pool are also automatically calculated and included in the report.
When used by radiology professionals, the standardized PYLARIFY AI report provides consistent and quantitative assessments of PSMA PET/CT.
PYLARIFY AI® Deployment and Integration to fit into the clinical workflow
PYLARIFY AI® deployment can be facilitated both as a secure web cloud application and as a local server application. The security and patient privacy are paramount to the installation of PYLARIFY AI®.
Once deployed, the adaptive application can be integrated to the institution image acquisition platforms or to the Picture Archiving and Communication Systems (PACS). The report generated from PYLARIFY AI®, in DICOM and PDF format, could be attached in the existing clinical impression for the physician’s review.
Seamless connection with existing clinical workflow (e.g., linking to picture archiving and communication system (PACS) delivers a unique combination of clinical utility in a workflow-friendly package.
PYLARIFY AI® is a deep-learning application
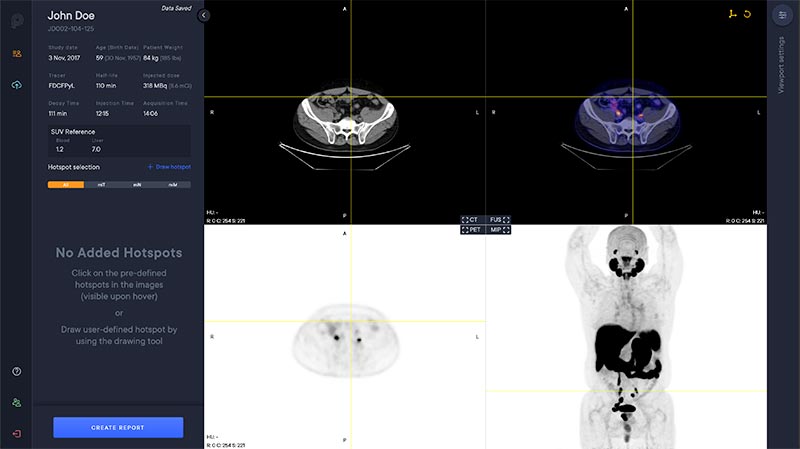
Viewer
The PYLARIFY AI viewer is a multi modal viewer with PET, CT, Fusion and MIP views. The available study information is shown in the control panel to the left.
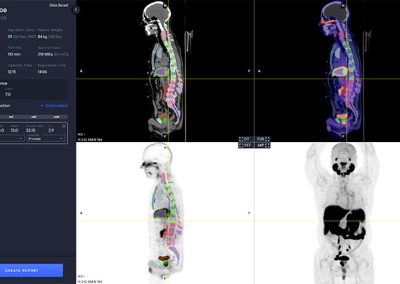
Segmentation
The software automatically analyzes the CT image to segment anatomical regions, including liver and aorta.
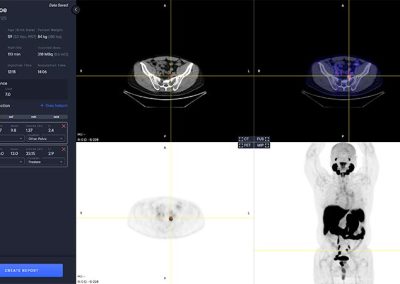
Hotspot Selection and Quantification
Hotspot selection is a tool that enables users to save volumes with high uptake for reporting and quantitative analysis. Each selected hotspot is shown in the left control panel.
References
- FDA clearance letter for aPROMISE X. Food and Drug Administration. April 29, 2022.
- Nickols N, et al. J Nucl Med. 2022;63(2):233-239.
- Johnsson K, et al. Eur J Nucl Med Mol Imaging. 2022;49(3):1041-1051.
- Jadvar H, et al. J Nucl Med. 2022;63(1):59-68.
- Hope TA, et al. J Nucl Med. 2023;64(9):1417.
- Kratochwil C, et al. Eur Jour Nuc Med Mol Imaging. 2023;50(9):2830-2845
- Calais J, et al. J NuclMed. 2022;63(supple2):2496
- Seifert R, et al. EurUrol. 2023;83(5):405-412.
- Kuo P, et al. J Clin Oncol. 2022;40(16 suppl):5002-5002
- ButeauJP, et al. Lancet Oncol. 2022;23(11):1389-1397.